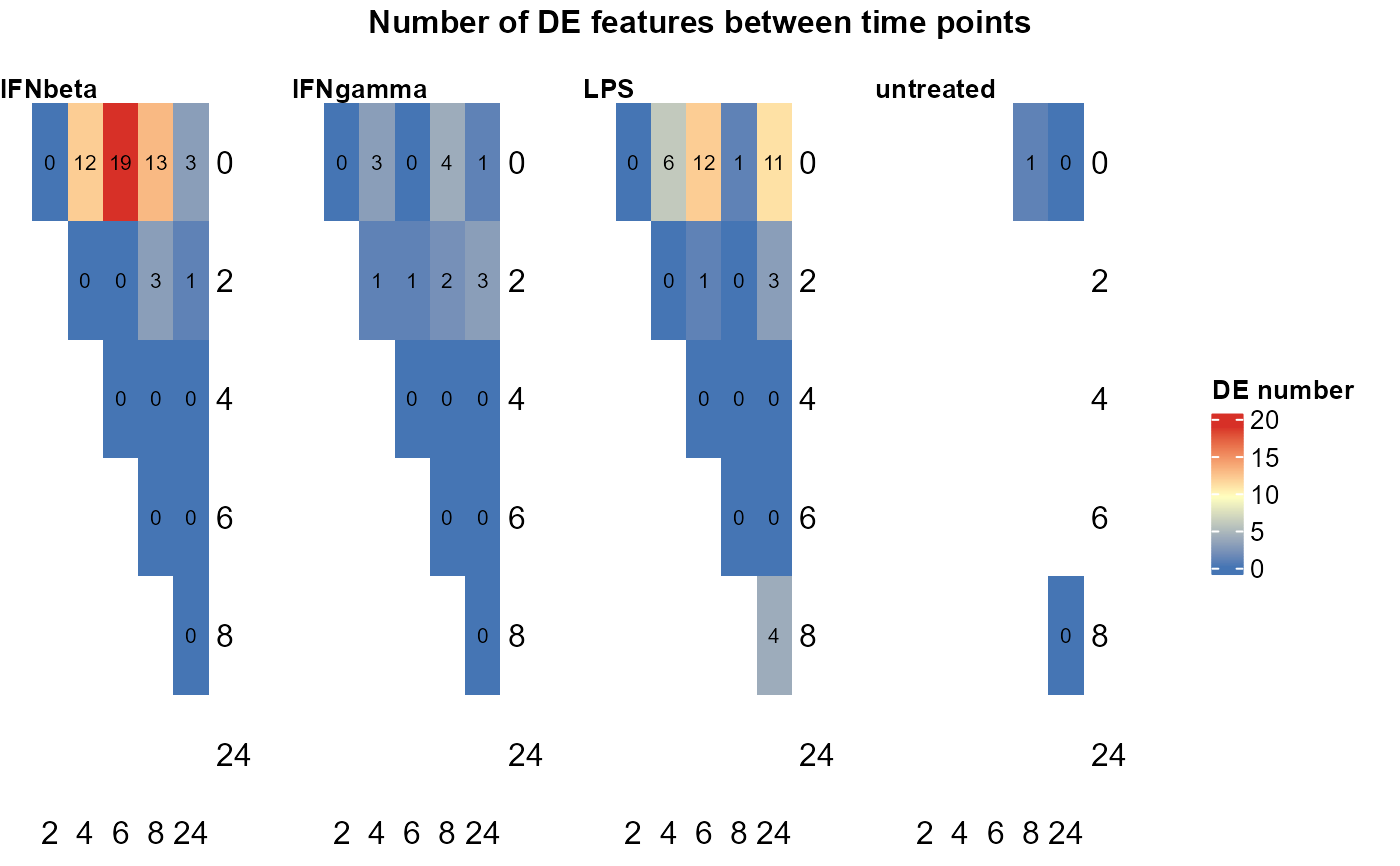
Differential expression analysis between time points within each group by limma
Usage
DE_between_time(
se_obj,
group = NULL,
filter = NULL,
assay = c(1, 2),
adjP_thres = 0.05,
logFC_thres = 1,
fontsize = 8
)Arguments
- se_obj
A SummarizedExperiment object created by
create_input- group
(Optional) A character vector specifying which groups to analyse. If NULL, all groups in the 'Group' column will be used. (default is NULL)
- filter
(Optional) Minimum number of replicates required in both conditions for a feature to be tested. If NULL, the minimum number of replicates across all groups and time points will be used. (default is NULL)
- assay
Assay index to use, where 1 is the original data and 2 is normalised to time 0 (if available) (default is 1)
- adjP_thres
(Optional) Threshold for adjusted p-value to consider a feature as differentially expressed (default is 0.05)
- logFC_thres
(Optional) Threshold for log2 fold change to consider a feature as differentially expressed (default is 1)
- fontsize
(Optional) Font size for the heatmap of DE numbers (default is 8)
Value
A list containing two elements: 'all_list' is a nested list of DE results for each group and time point comparison including all features; 'de_list' is a nested list of filtered DE results based on the specified thresholds including only significant features.
Examples
data("example")
example_obj <- normalise_to_start(example_obj)
DE_between_time_out <- DE_between_time(example_obj, assay = 1)
#> Non-NA replicate number filter not specified. Using minimum number of replicates across all groups and time points: 0
#> Comparing group IFNbeta time 2 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNbeta time 4 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNbeta time 6 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNbeta time 8 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNbeta time 24 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNbeta time 4 to 2: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNbeta time 6 to 2: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNbeta time 8 to 2: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNbeta time 24 to 2: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNbeta time 6 to 4: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNbeta time 8 to 4: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNbeta time 24 to 4: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNbeta time 8 to 6: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNbeta time 24 to 6: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNbeta time 24 to 8: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNgamma time 2 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNgamma time 4 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNgamma time 6 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNgamma time 8 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNgamma time 24 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNgamma time 4 to 2: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNgamma time 6 to 2: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNgamma time 8 to 2: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNgamma time 24 to 2: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNgamma time 6 to 4: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNgamma time 8 to 4: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNgamma time 24 to 4: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNgamma time 8 to 6: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNgamma time 24 to 6: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group IFNgamma time 24 to 8: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group LPS time 2 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group LPS time 4 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group LPS time 6 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group LPS time 8 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group LPS time 24 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group LPS time 4 to 2: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group LPS time 6 to 2: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group LPS time 8 to 2: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group LPS time 24 to 2: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group LPS time 6 to 4: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group LPS time 8 to 4: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group LPS time 24 to 4: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group LPS time 8 to 6: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group LPS time 24 to 6: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group LPS time 24 to 8: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group untreated time 8 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group untreated time 24 to 0: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
#> Comparing group untreated time 24 to 8: keeping 100 of 100 features (100.0%)
#> Warning: Zero sample variances detected, have been offset away from zero
plot_DE_between_time(example_obj,
de_list = DE_between_time_out$de_list,
fontsize = 8, value = TRUE, nrow = 1, heatmap_width = 3)
#> Warning: The input is a data frame-like object, convert it to a matrix.
#> Warning: The input is a data frame-like object, convert it to a matrix.
#> Warning: The input is a data frame-like object, convert it to a matrix.
#> Warning: The input is a data frame-like object, convert it to a matrix.
#> Warning: Note: not all columns in the data frame are numeric. The data frame
#> will be converted into a character matrix.
#> Registered S3 method overwritten by 'car':
#> method from
#> na.action.merMod lme4