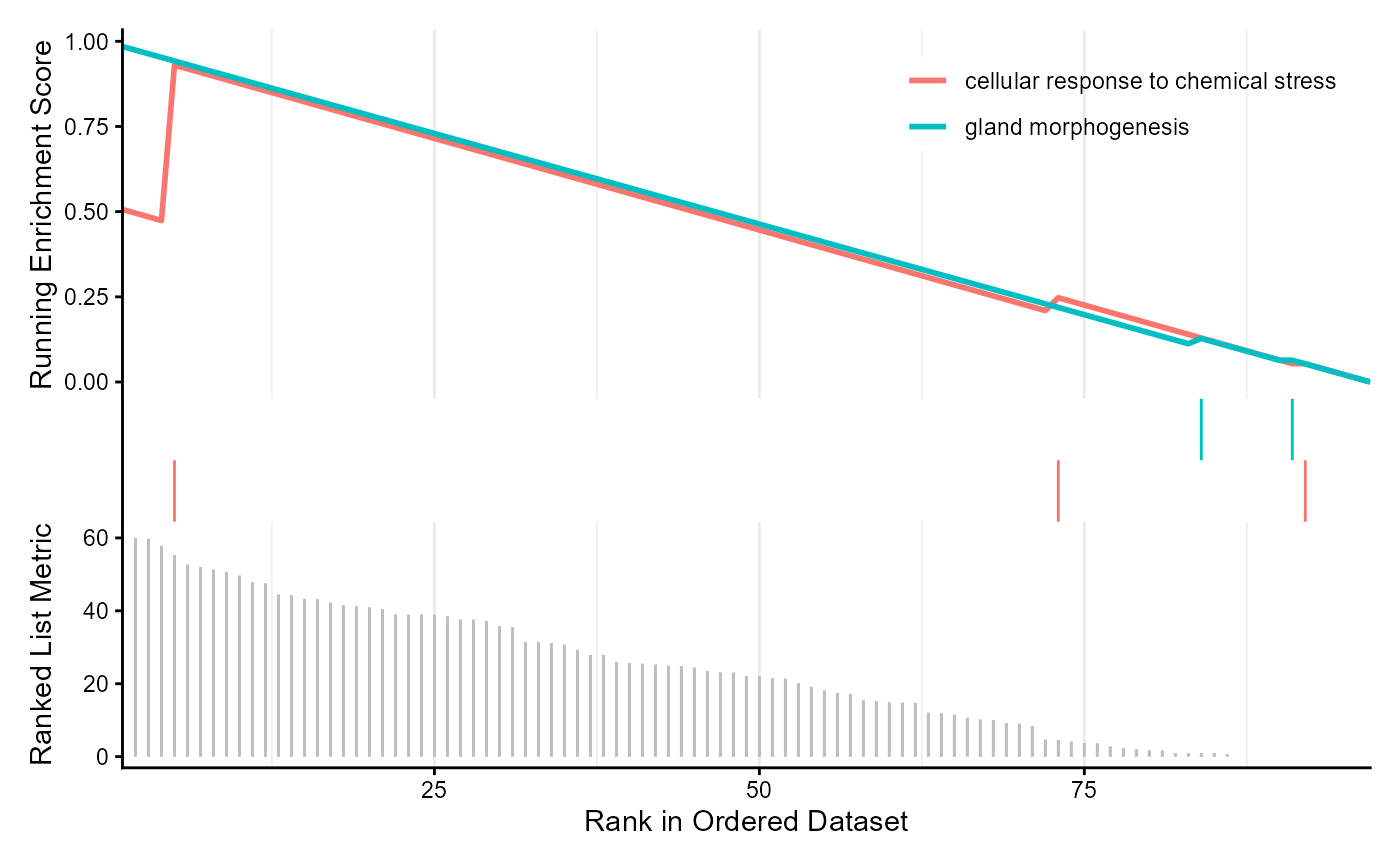
Gene ontology enrichment analysis of a ranked gene list with clusterProfiler. Genes can be ranked by variance decomposition results.
Usage
enrichGO_rank(
rank_table,
gene_rank_by,
OrgDb,
keyType = "SYMBOL",
go_rank_by = "p.adjust",
category = NULL,
pvalueCutoff = 0.05,
pAdjustMethod = "BH",
...
)Arguments
- rank_table
A data frame with at least two columns: 'Feature' for gene names and one or more columns of variables for ranking the genes, e.g. output of
decomp_variance()containing variance decomposition results- gene_rank_by
Variable in
rank_tableto rank the genes by, e.g. "Time", "Group" in the output ofdecomp_variance()- OrgDb
Organism database, e.g. org.Hs.eg.db, org.Mm.eg.db
- keyType
(Optional) Available options are
AnnotationDbi::keytypes(OrgDb)(default is "SYMBOL")- go_rank_by
(Optional) Variable in the GO enrichment result to rank the GO terms by (default is "p.adjust", other options include "pvalue", "qvalue", "NES", "setSize", "enrichmentScore", etc.)
- category
(Optional) GO category to analyze (default is all three of BP, MF, CC)
- pvalueCutoff
(Optional) Parameter of
clusterProfiler::gseGO()(default is 0.05)- pAdjustMethod
(Optional) Parameter of
clusterProfiler::gseGO()(default is "BH")- ...
additional arguments passed to
clusterProfiler::gseGO()
Examples
library(org.Mm.eg.db)
data("example")
example_obj <- normalise_to_start(example_obj)
var_decomp <- decomp_variance(example_obj, assay = 1)
#>
| | 0 % ~calculating
|+ | 1 % ~01s
|+ | 2 % ~01s
|++ | 3 % ~01s
|++ | 4 % ~01s
|+++ | 5 % ~01s
|+++ | 6 % ~01s
|++++ | 7 % ~01s
|++++ | 8 % ~01s
|+++++ | 9 % ~01s
|+++++ | 10% ~01s
|++++++ | 11% ~01s
|++++++ | 12% ~01s
|+++++++ | 13% ~01s
|+++++++ | 14% ~01s
|++++++++ | 15% ~01s
|++++++++ | 16% ~01s
|+++++++++ | 17% ~01s
|+++++++++ | 18% ~01s
|++++++++++ | 19% ~01s
|++++++++++ | 20% ~01s
|+++++++++++ | 21% ~01s
|+++++++++++ | 22% ~01s
|++++++++++++ | 23% ~01s
|++++++++++++ | 24% ~01s
|+++++++++++++ | 25% ~01s
|+++++++++++++ | 26% ~01s
|++++++++++++++ | 27% ~01s
|++++++++++++++ | 28% ~01s
|+++++++++++++++ | 29% ~01s
|+++++++++++++++ | 30% ~01s
|++++++++++++++++ | 31% ~01s
|++++++++++++++++ | 32% ~01s
|+++++++++++++++++ | 33% ~01s
|+++++++++++++++++ | 34% ~01s
|++++++++++++++++++ | 35% ~01s
|++++++++++++++++++ | 36% ~01s
|+++++++++++++++++++ | 37% ~01s
|+++++++++++++++++++ | 38% ~01s
|++++++++++++++++++++ | 39% ~01s
|++++++++++++++++++++ | 40% ~01s
|+++++++++++++++++++++ | 41% ~01s
|+++++++++++++++++++++ | 42% ~01s
|++++++++++++++++++++++ | 43% ~01s
|++++++++++++++++++++++ | 44% ~01s
|+++++++++++++++++++++++ | 45% ~01s
|+++++++++++++++++++++++ | 46% ~01s
|++++++++++++++++++++++++ | 47% ~01s
|++++++++++++++++++++++++ | 48% ~01s
|+++++++++++++++++++++++++ | 49% ~01s
|+++++++++++++++++++++++++ | 50% ~01s
|++++++++++++++++++++++++++ | 51% ~01s
|++++++++++++++++++++++++++ | 52% ~01s
|+++++++++++++++++++++++++++ | 53% ~01s
|+++++++++++++++++++++++++++ | 54% ~01s
|++++++++++++++++++++++++++++ | 55% ~01s
|++++++++++++++++++++++++++++ | 56% ~01s
|+++++++++++++++++++++++++++++ | 57% ~01s
|+++++++++++++++++++++++++++++ | 58% ~01s
|++++++++++++++++++++++++++++++ | 59% ~01s
|++++++++++++++++++++++++++++++ | 60% ~00s
|+++++++++++++++++++++++++++++++ | 61% ~00s
|+++++++++++++++++++++++++++++++ | 62% ~00s
|++++++++++++++++++++++++++++++++ | 63% ~00s
|++++++++++++++++++++++++++++++++ | 64% ~00s
|+++++++++++++++++++++++++++++++++ | 65% ~00s
|+++++++++++++++++++++++++++++++++ | 66% ~00s
|++++++++++++++++++++++++++++++++++ | 67% ~00s
|++++++++++++++++++++++++++++++++++ | 68% ~00s
|+++++++++++++++++++++++++++++++++++ | 69% ~00s
|+++++++++++++++++++++++++++++++++++ | 70% ~00s
|++++++++++++++++++++++++++++++++++++ | 71% ~00s
|++++++++++++++++++++++++++++++++++++ | 72% ~00s
|+++++++++++++++++++++++++++++++++++++ | 73% ~00s
|+++++++++++++++++++++++++++++++++++++ | 74% ~00s
|++++++++++++++++++++++++++++++++++++++ | 75% ~00s
|++++++++++++++++++++++++++++++++++++++ | 76% ~00s
|+++++++++++++++++++++++++++++++++++++++ | 77% ~00s
|+++++++++++++++++++++++++++++++++++++++ | 78% ~00s
|++++++++++++++++++++++++++++++++++++++++ | 79% ~00s
|++++++++++++++++++++++++++++++++++++++++ | 80% ~00s
|+++++++++++++++++++++++++++++++++++++++++ | 81% ~00s
|+++++++++++++++++++++++++++++++++++++++++ | 82% ~00s
|++++++++++++++++++++++++++++++++++++++++++ | 83% ~00s
|++++++++++++++++++++++++++++++++++++++++++ | 84% ~00s
|+++++++++++++++++++++++++++++++++++++++++++ | 85% ~00s
|+++++++++++++++++++++++++++++++++++++++++++ | 86% ~00s
|++++++++++++++++++++++++++++++++++++++++++++ | 87% ~00s
|++++++++++++++++++++++++++++++++++++++++++++ | 88% ~00s
|+++++++++++++++++++++++++++++++++++++++++++++ | 89% ~00s
|+++++++++++++++++++++++++++++++++++++++++++++ | 90% ~00s
|++++++++++++++++++++++++++++++++++++++++++++++ | 91% ~00s
|++++++++++++++++++++++++++++++++++++++++++++++ | 92% ~00s
|+++++++++++++++++++++++++++++++++++++++++++++++ | 93% ~00s
|+++++++++++++++++++++++++++++++++++++++++++++++ | 94% ~00s
|++++++++++++++++++++++++++++++++++++++++++++++++ | 95% ~00s
|++++++++++++++++++++++++++++++++++++++++++++++++ | 96% ~00s
|+++++++++++++++++++++++++++++++++++++++++++++++++ | 97% ~00s
|+++++++++++++++++++++++++++++++++++++++++++++++++ | 98% ~00s
|++++++++++++++++++++++++++++++++++++++++++++++++++| 99% ~00s
|++++++++++++++++++++++++++++++++++++++++++++++++++| 100% elapsed=01s
example_go_rank <- enrichGO_rank(var_decomp, gene_rank_by = "Time",
OrgDb = org.Mm.eg.db, keyType = "SYMBOL", category = "BP")
#> NA / NaN / Inf values found in the gene ranking variable. Those genes will be removed.
#> Warning: There are ties in the preranked stats (1.03% of the list). The order of those tied genes will be arbitrary, which may produce unexpected results.
#> Warning: There were 4821 pathways for which P-values were not calculated properly due to unbalanced gene-level statistic values. For such pathways pvalue, NES and log2err are set to NA. You can try to increase nPermSimple.
#> Warning: Invalid p-values detected (NA, non-finite, <0, or >1). qvalue will be computed on valid p-values only.
#> Warning: NA values detected in gene set IDs. Replacing with string 'NA'.
#> Warning: Duplicate gene set IDs detected: NA... (Total 1). Unique suffixes added.
#> Removing NA ID gene sets.
enrichplot::gseaplot2(example_go_rank, geneSetID = 1:2)