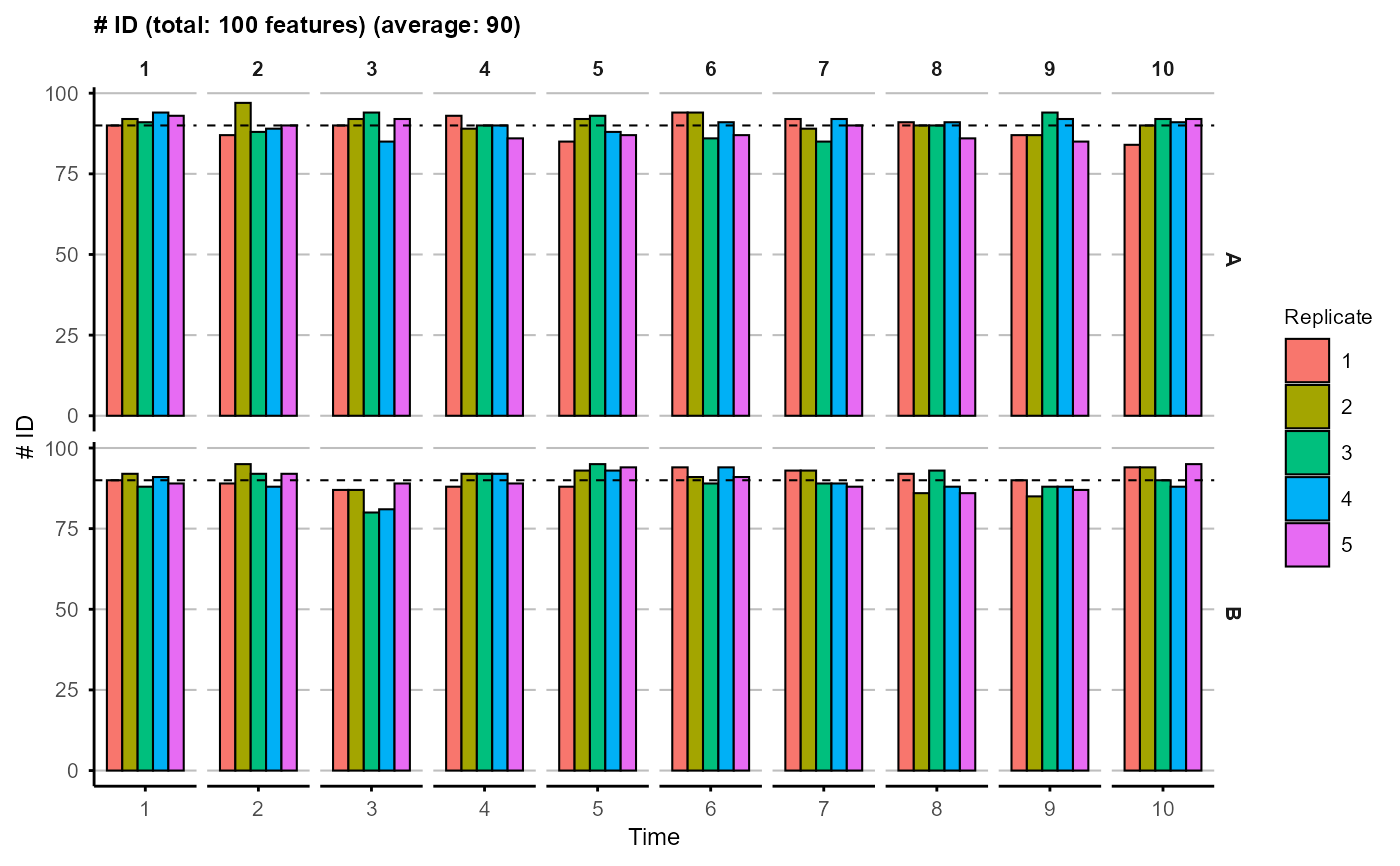
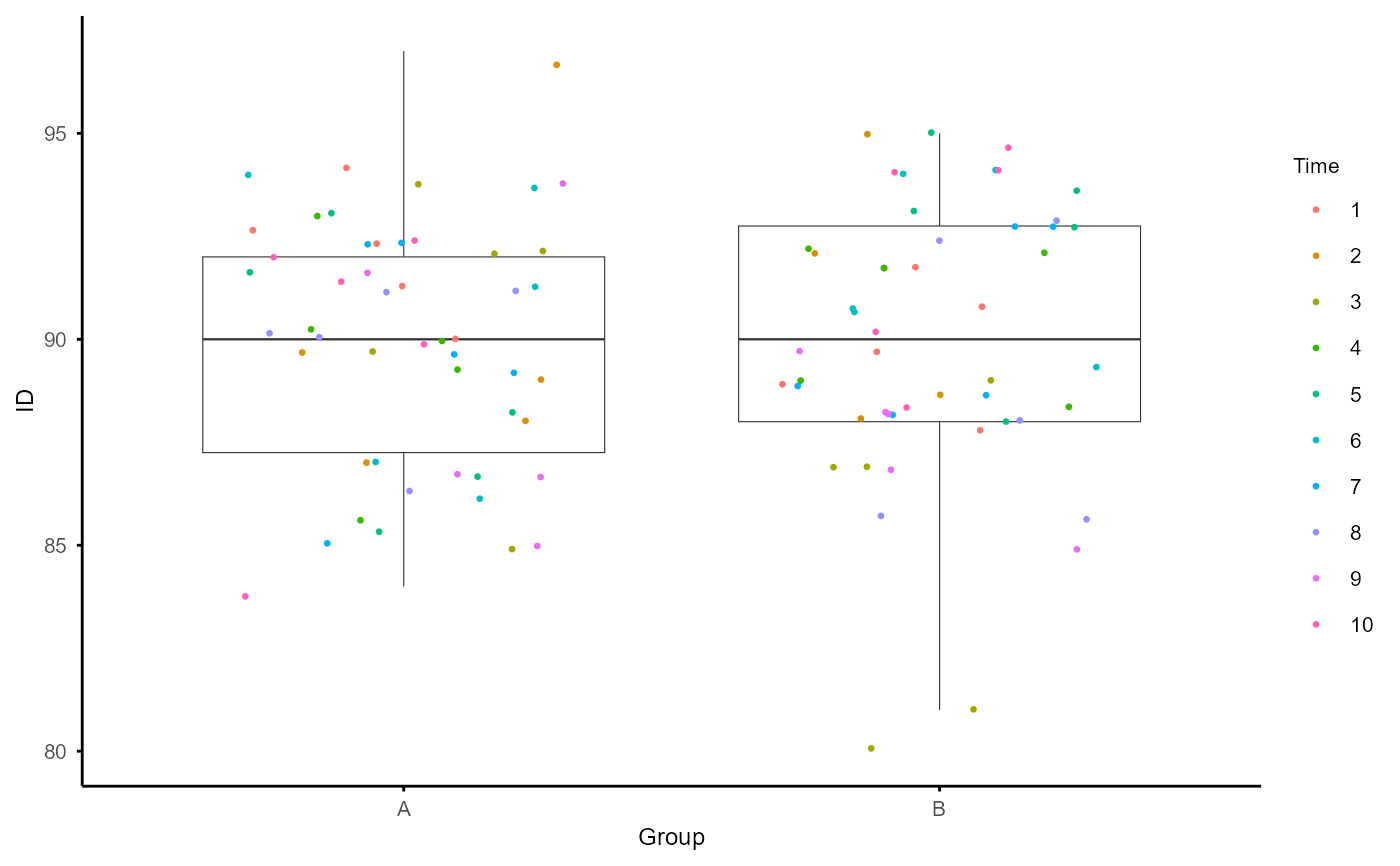
Plot the number of identified features (i.e., features with non-missing values) for each sample, with a dashed line indicating the average number across all samples.
Arguments
- se_obj
A SummarizedExperiment object, produced by
create_input()function, containing the abundance data and associated sample information.- fontsize
(Optional) An integer specifying the font size for the plot (default is 8).
- signif
(Optional) A logical value indicating whether to perform significance testing between groups and add significance annotations to the plot (default is FALSE).
- ...
Additional arguments to be passed to
ggsignif::geom_signif()function whensignifis TRUE, for customizing the significance annotations.
Value
A plot showing the ID number for each sample, with a dashed line indicating the average number across all samples.
Examples
# simulate data with random missing values
na_data <- matrix(rnorm(1000), nrow = 100, ncol = 100)
na_data[sample(length(na_data), size = 1000)] <- NA
na_data <- data.frame(Feature = paste0("Feature", 1:100), na_data)
colnames(na_data)[-1] <- paste0("Sample", 1:100)
na_obj <- create_input(na_data,
data.frame(Sample = paste0("Sample", 1:100),
Time = rep(rep(1:10, each = 5)), 2,
Group = rep(c("A", "B"), each = 50),
Replicate = rep(1:5, 20)))
#> Converting 'Group' column to factor. Default order is alphabetical.
#> Converting 'Replicate' column to factor. Default order is numerical.
plot_ID(na_obj)
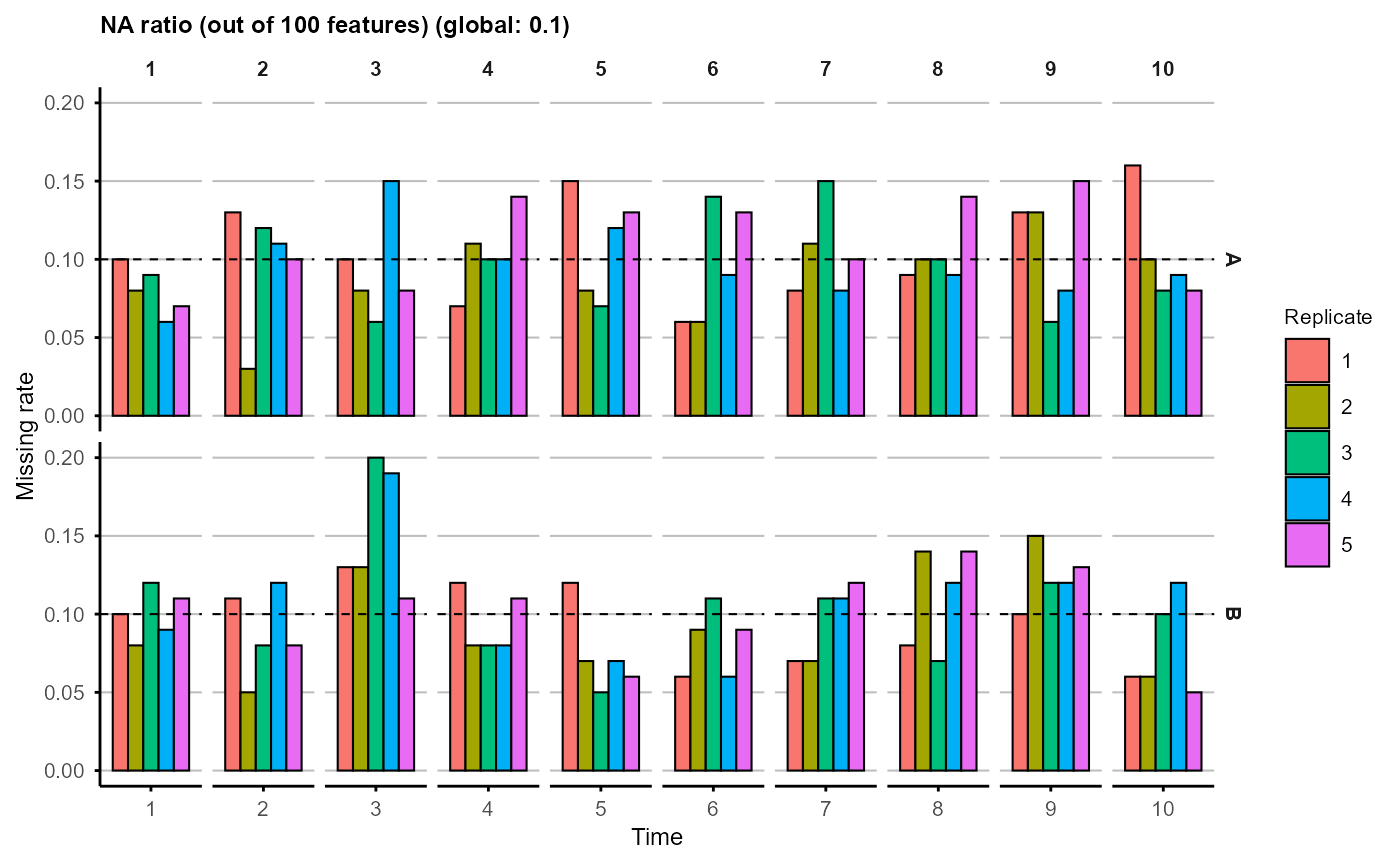
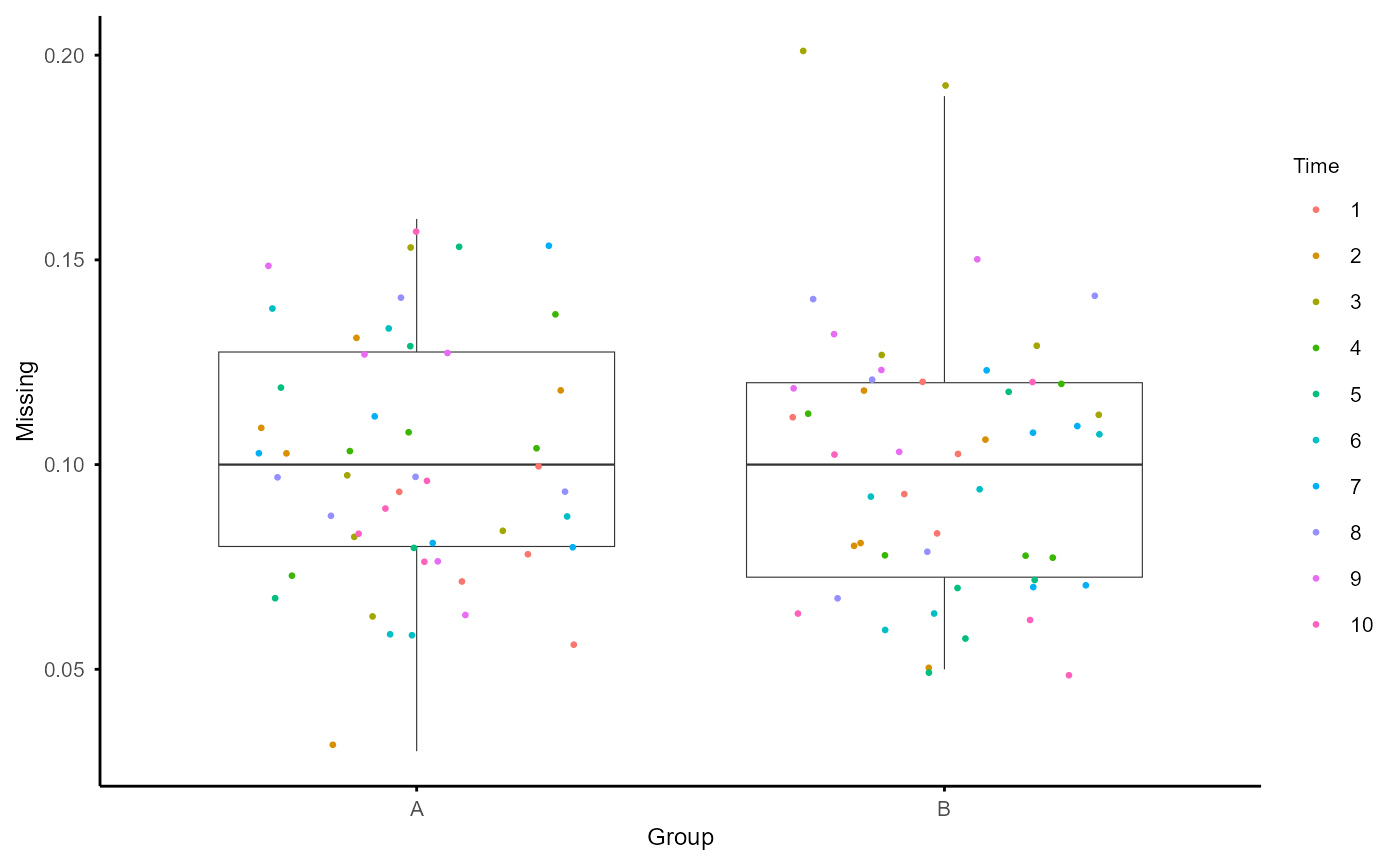
 plot_missing(na_obj)
plot_missing(na_obj)