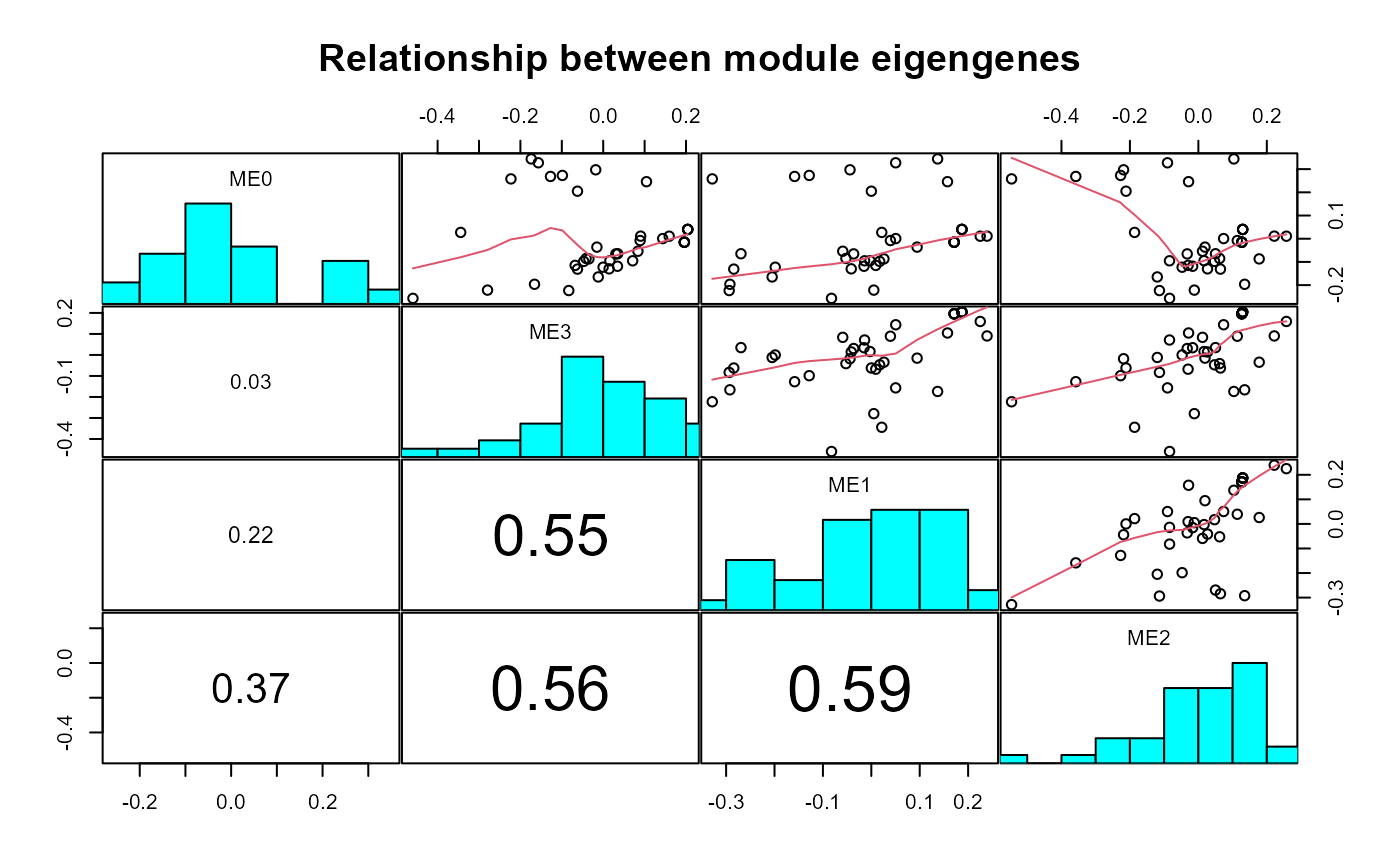
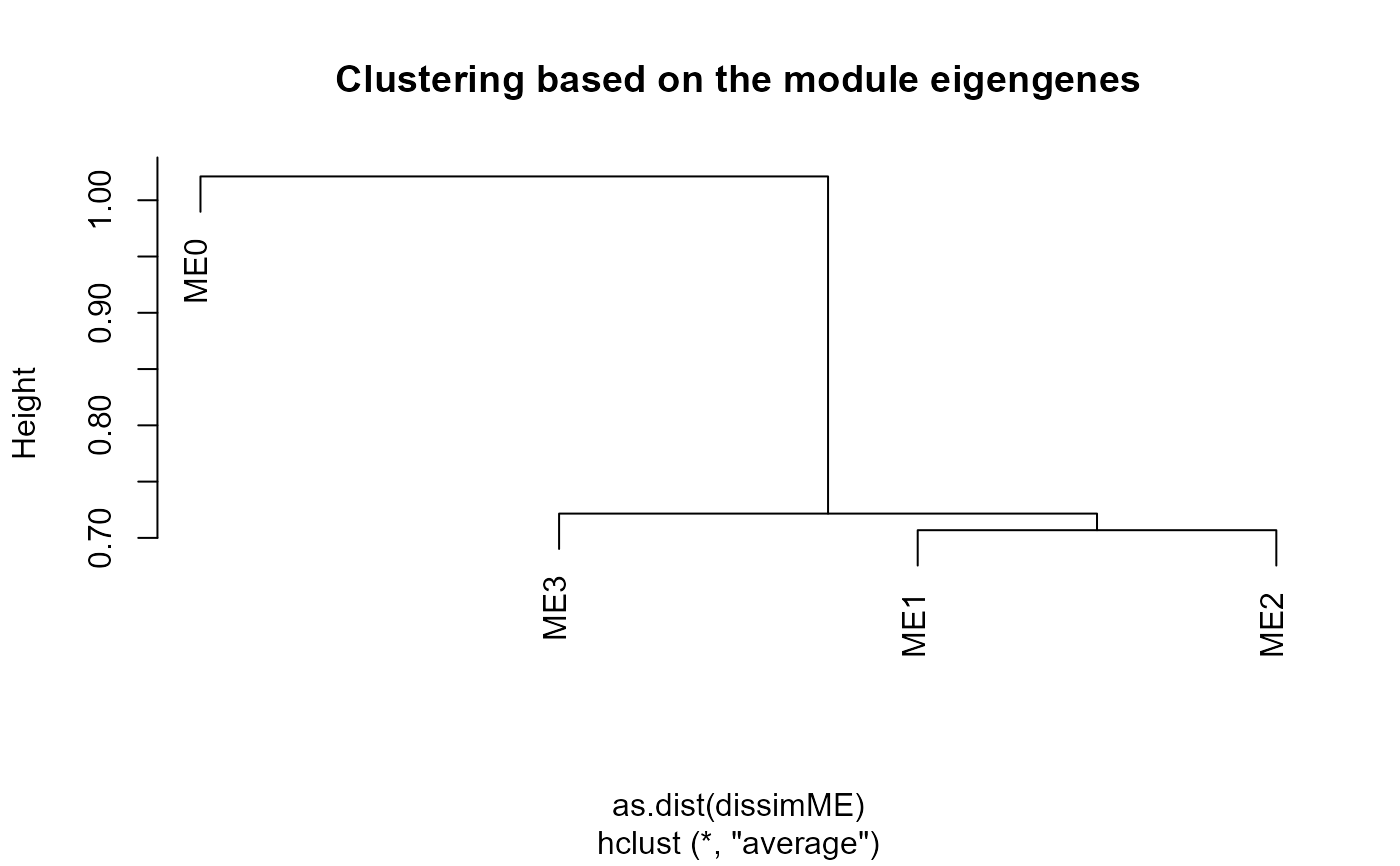
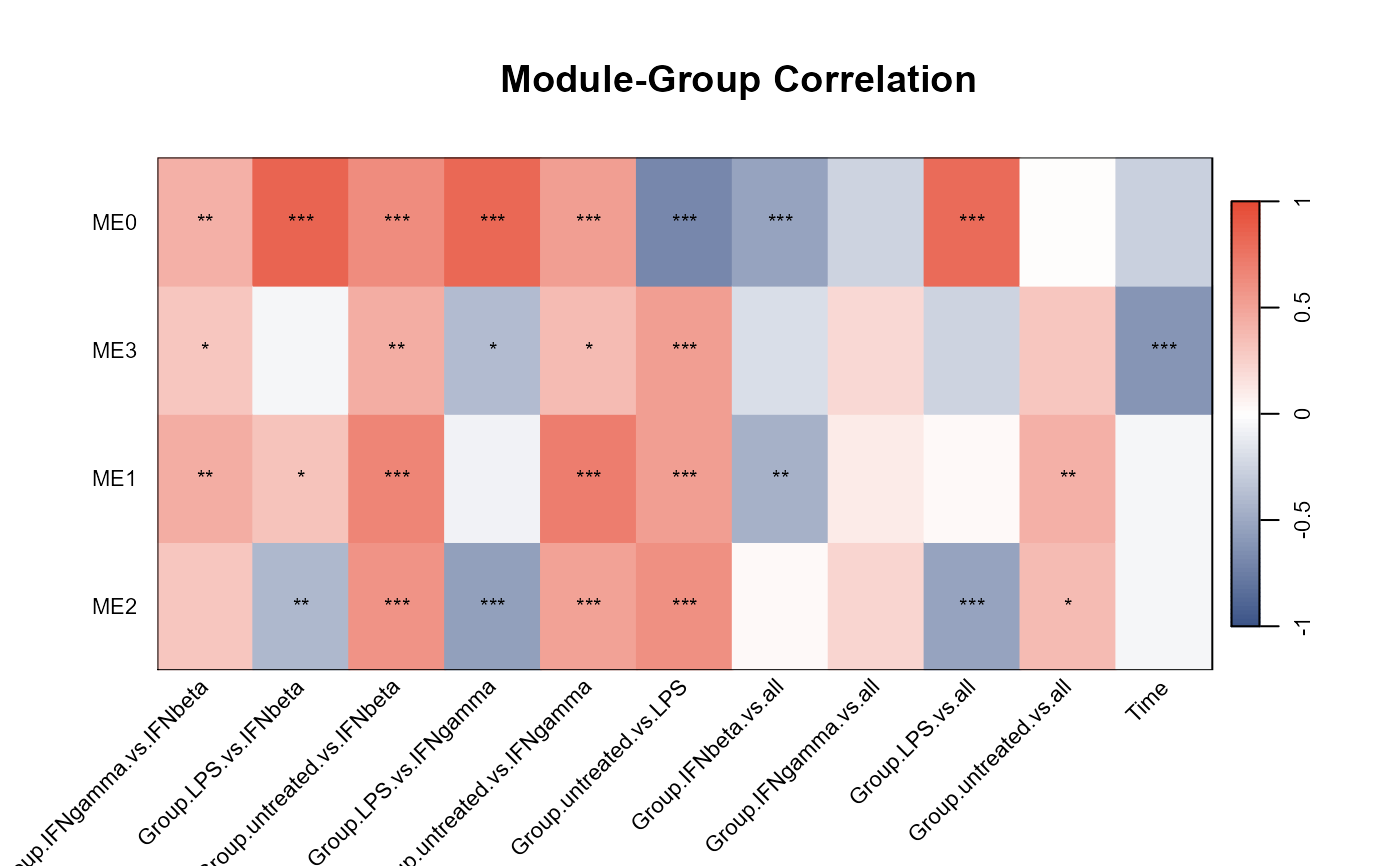
Plot WGCNA module eigengenes and module-group/time correlations
based on output of run_WGCNA()
Arguments
- net
WGCNA network object output by
run_WGCNA()- fontsize
Font size for plots (default is 8)
Value
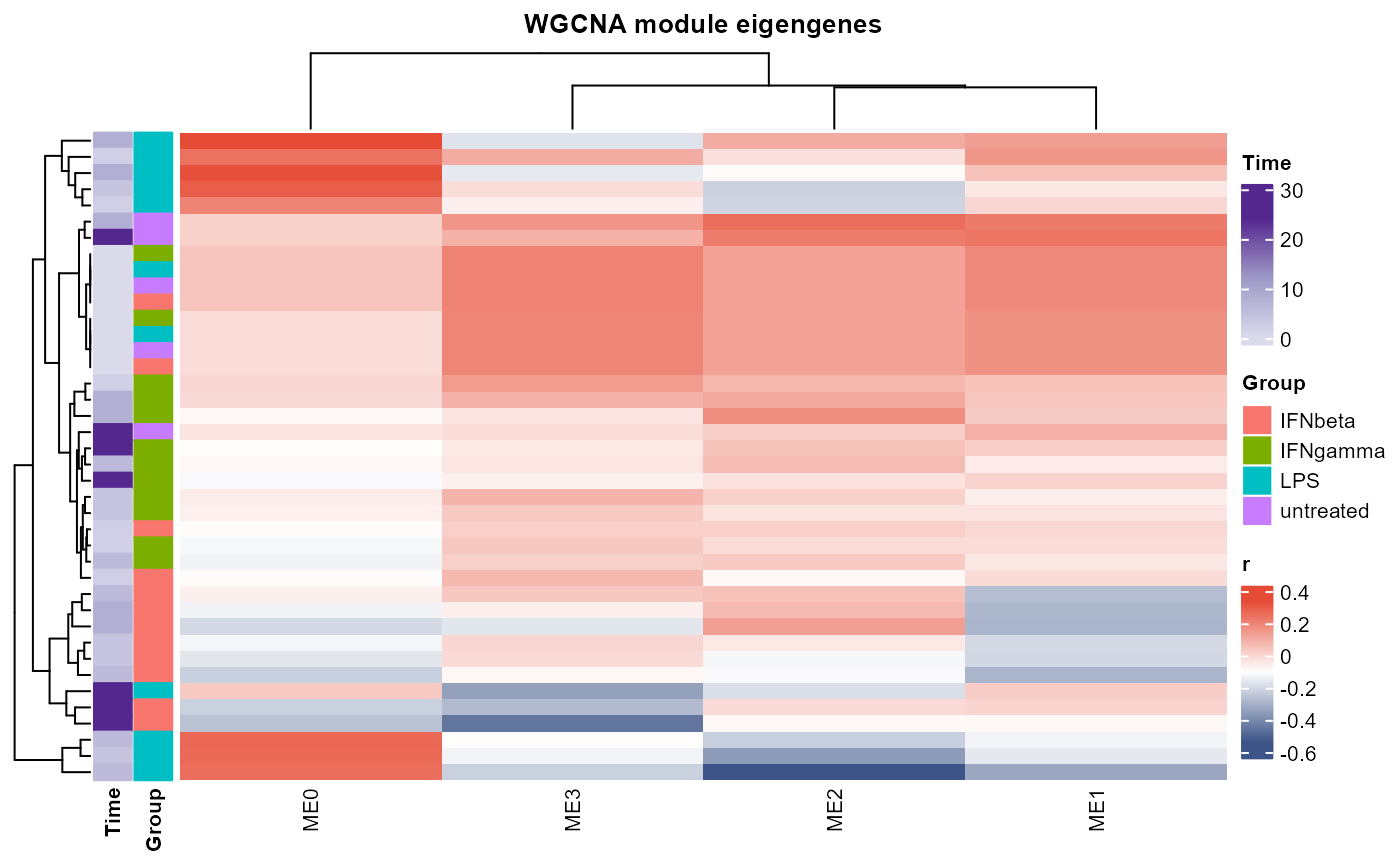
Plots of WGCNA module dendrogram, module eigengenes, pairwise scatterplots of eigengenes, clustering of module eigengenes, and module-trait correlation heatmap
References
https://github.com/edo98811/WGCNA_official_documentation/blob/main/FemaleLiver-03-relateModsToExt.R
Examples
data("example")
example_obj <- normalise_to_start(example_obj)
# wgcna_input <- prepare_WGCNA(example_obj, assay = 2, powers = seq(1, 30),
# networkType = "signed", RsquaredCut = 0.8)
# wgcna_input$fitIndices
# picked_power <- wgcna_input$powerEstimate
# example_net <- run_WGCNA(wgcna_input,
# power = picked_power,
# minModuleSize = 10, # only 100 genes in the example data
# numericLabels = TRUE)
data("example_net")
plot_WGCNA(example_net, fontsize = 8)
#> Warning: The input is a data frame-like object, convert it to a matrix.
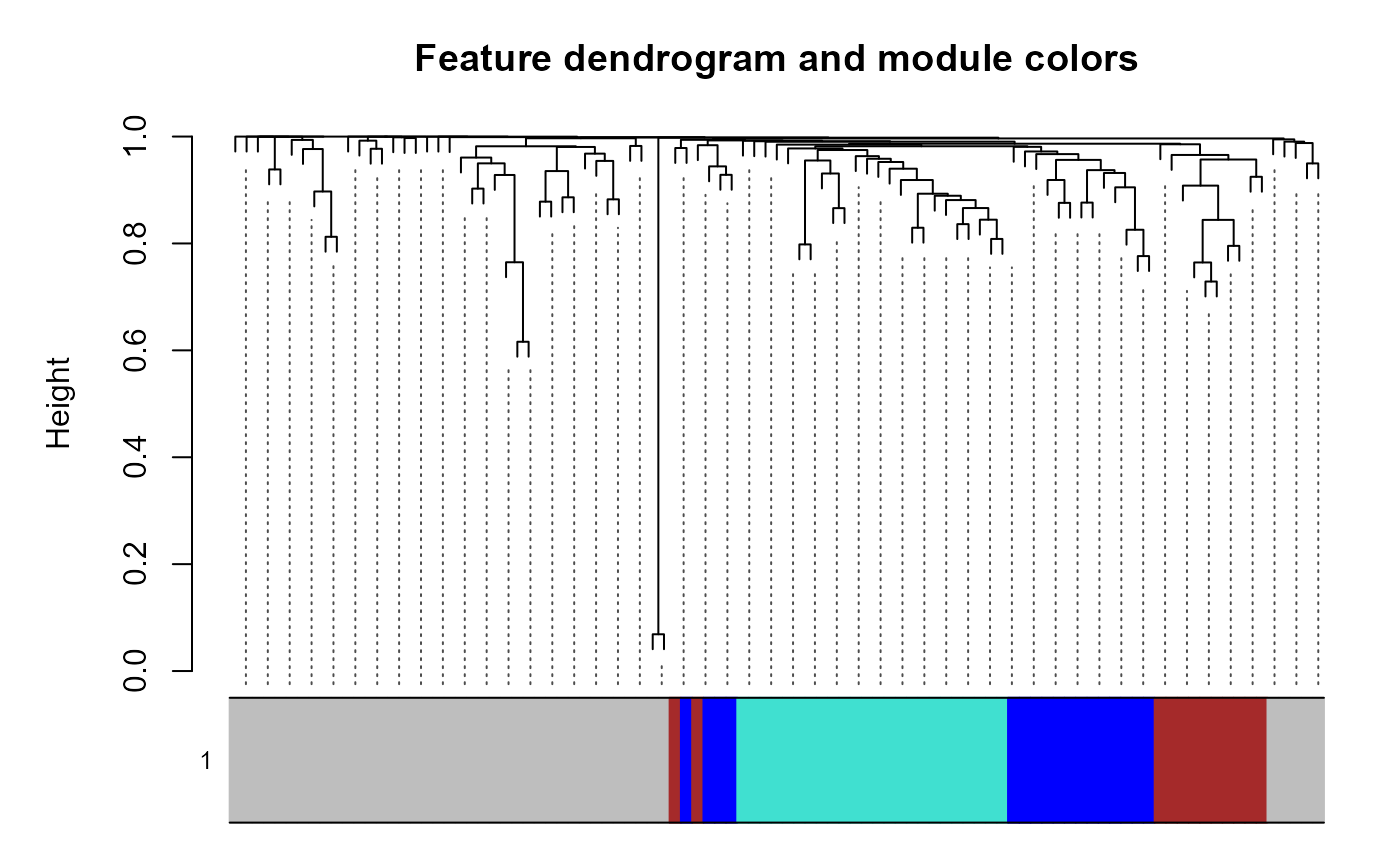
 #> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter
#> Warning: argument 1 does not name a graphical parameter