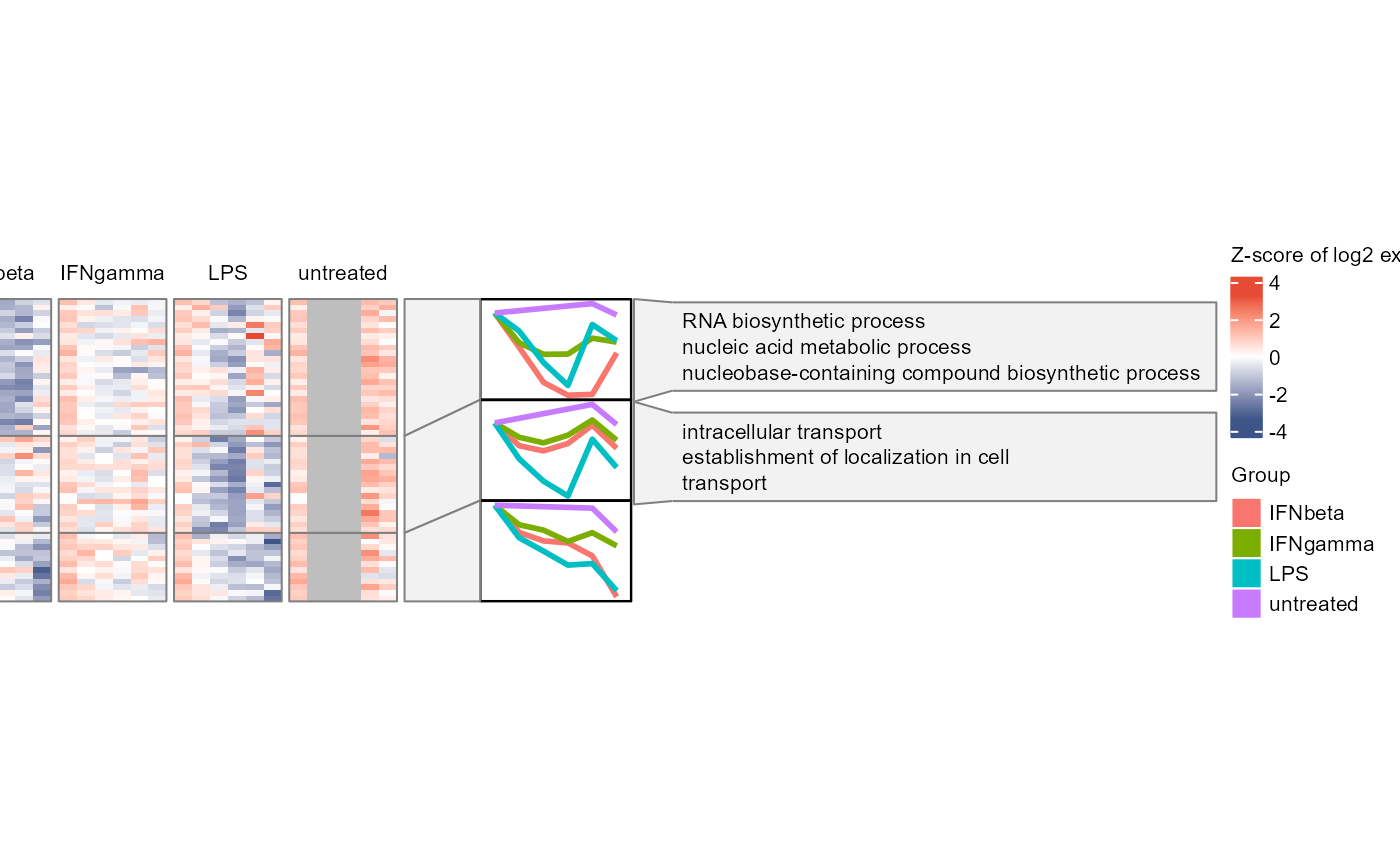
Plot WGCNA modules' heatmaps, mean expression profiles, and GO terms, align horizontally.
Different from running WGCNA, the input data should have the replicates merged, instead of having multiple samples per group, time and feature (gene).
If certain time points are missing in some groups, NA values are added.
Usage
plot_modules_h(
module,
se_obj_merged,
scale = TRUE,
assay = 2,
ylabel = "Log2 abundance",
suffix = "",
device = c("png", "pdf", "tiff", "jpeg"),
save = NULL,
profile_width = 3,
profile_link_width = 1,
go_list = NULL,
go_category = "BP",
go_rank_by = "p.adjust",
go_top_n = 3,
fontsize = 8,
heatmap_width = 8,
heatmap_height = 8,
width = 16,
height = 12,
res = 300
)Arguments
- module
A data frame with columns "Feature" and "Module"
- se_obj_merged
A SummarizedExperiment object, with one value for each feature at each time point in each group (replicates merged). The colData of the object should contain columns "Sample", "Group", and "Time". The object can be produced by
split_groups(),merge_replicates()andmerge_group().- scale
Whether to scale the data (z-score) across samples for each feature (default is TRUE)
- assay
The assay index in the SummarizedExperiment object to use (default is 2, time 0 normalised data)
- ylabel
Y axis label prefix (default is "Abundance")
- suffix
Suffix for the saved image file name (default is an empty string)
- device
Image file format for saving (default is "png")
- save
Directory to save the plot, no saving if is NULL (default is NULL)
- profile_width
Width of the mean expression profile plot (default is 3 (cm))
- profile_link_width
Width of the link in cm between heatmap and mean expression profile plot (default is 1)
- go_list
A list of GO enrichment results for WGCNA modules, as produced by the
enrichGO_list()function. If NULL (default), no GO terms will be displayed.- go_category
Category of GO terms to display, one of "BP", "MF", or "CC" (default is "BP")
- go_rank_by
Column name in the GO enrichment result to rank the GO terms (default is "p.adjust")
- go_top_n
Number of top GO terms to display for each module (default is 3)
- fontsize
Font size (default is 8)
- heatmap_width
Width of the heatmap body in cm (default is 8)
- heatmap_height
Height of the heatmap body in cm (default is 8)
- width
Width of the saved image (default is 16 (cm))
- height
Height of the saved image (default is 12 (cm))
- res
Resolution of the saved image (except pdf format) (default is 300 (ppi))
Examples
library(dplyr)
library(org.Mm.eg.db)
data(example)
example_obj <- normalise_to_start(example_obj)
example_obj_list <- split_groups(example_obj)
example_obj_merged_list <- merge_replicates(example_obj_list)
example_obj_merged <- merge_groups(example_obj_merged_list)
data(example_net)
example_module <- data.frame(Module = as.factor(example_net$colors)) %>%
tibble::rownames_to_column("Feature") %>% arrange(Module)
# select two modules for demonstration
example_module_list <- example_module %>%
filter(Module %in% c(1, 2)) %>%
split(as.character(.$Module)) %>%
lapply(`[[`, "Feature")
# set cutoff to 1 to show all results for demonstration
example_go_list = enrichGO_list(example_module_list, OrgDb = org.Mm.eg.db,
universe = example_module$Feature,
pvalueCutoff = 1, qvalueCutoff = 1,
category = "BP", simplify = FALSE)
#> Performing GO enrichment for category: BP
#> Processing gene list: 1
#> 'select()' returned 1:1 mapping between keys and columns
#> 'select()' returned 1:1 mapping between keys and columns
#> Warning: 1.03% of input gene IDs are fail to map...
#> Processing gene list: 2
#> 'select()' returned 1:1 mapping between keys and columns
#> Warning: 5.88% of input gene IDs are fail to map...
#> 'select()' returned 1:1 mapping between keys and columns
#> Warning: 1.03% of input gene IDs are fail to map...
#> Merging GO enrichment results across gene lists for each category.
# plot_GO(example_go_list$all, plot_dotplot = TRUE,
# plot_emapplot = FALSE, plot_cnetplot = FALSE)
plot_modules_h(example_module %>% filter(Module != '0'),
example_obj_merged, scale = TRUE,
ylabel = "Z-score of log2 expression",
go_list = example_go_list$all, go_category = "BP",
heatmap_width = 6, heatmap_height = 4)
#> Warning: Removed 72 rows containing non-finite outside the scale range
#> (`stat_summary()`).
#> Warning: Removed 51 rows containing non-finite outside the scale range
#> (`stat_summary()`).
#> Warning: Removed 36 rows containing non-finite outside the scale range
#> (`stat_summary()`).