UMAP
=======
Plot UMAP
>>>>>>> Stashed changes Source:R/plot_umap.R
plot_umap.RdPlot UMAP of samples, using features without missing values
The UMAP plots are labelled by Group, Time, Replicate (if more than 1), Batch (if more than 1), and number of identified features (non-missing values).
Usage
plot_umap(
se_obj,
seed = 1234,
plot = TRUE,
plot_ID = FALSE,
circle = FALSE,
xlim_min = NULL,
xlim_max = NULL,
ylim_min = NULL,
ylim_max = NULL,
umap_neighbors = NULL,
fontsize = 8,
assay = 1,
...
)Arguments
- se_obj
A SummarizedExperiment object created by
create_input()- seed
Random seed for UMAP (default is 1234)
- plot
Whether to plot the figures (default is TRUE)
- plot_ID
Whether to include a UMAP plot coloured by number of identified features (default is FALSE)
- circle
Logical, whether to draw circles (ellipses) around samples of each group (default is FALSE)
- xlim_min
Minimum x-axis limit when drawing ellipses (default 1.5*min UMAP x)
- xlim_max
Maximum x-axis limit when drawing ellipses (default 1.5*max UMAP x)
- ylim_min
Minimum y-axis limit when drawing ellipses (default 1.5*min UMAP y)
- ylim_max
Maximum y-axis limit when drawing ellipses (default 1.5*max UMAP y)
- umap_neighbors
UMAP n_neighbors parameter (default is selected by
.umap_n_neighbors()function based on the number of samples)- fontsize
Font size for the plot (default is 8)
- assay
Assay index to use, where 1 is the original data and 2 is normalised to time 0 (if available) (default is 1)
- ...
Additional arguments passed to
ggforce::geom_mark_ellipse()for customizing the ellipses
Value
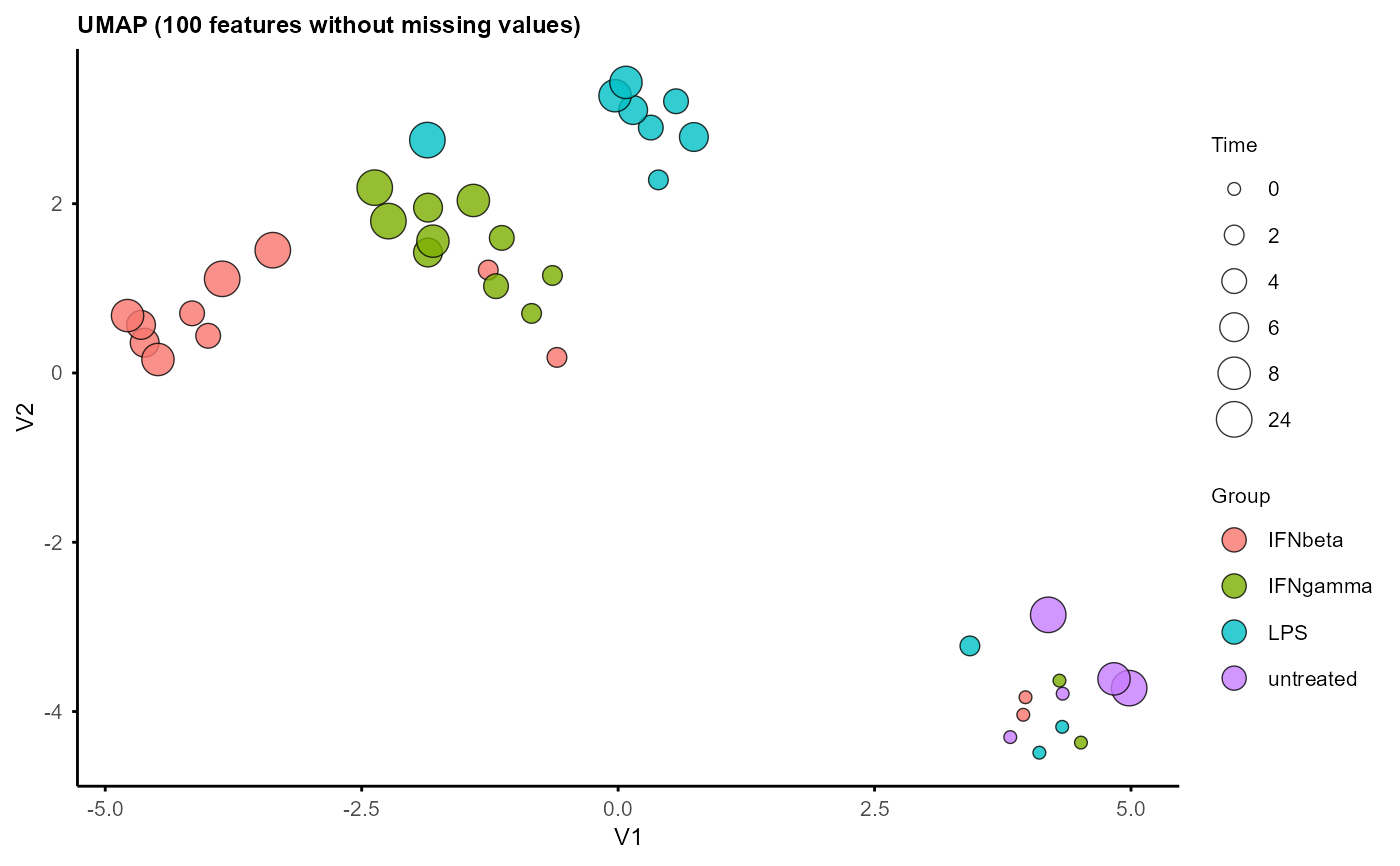
A UMAP plot showing the distribution of samples. And a data frame containing UMAP coordinates and sample annotations for custom plotting.
Examples
data("example")
umap_layout <- plot_umap(example_obj)
#> Using n_neighbors = 8
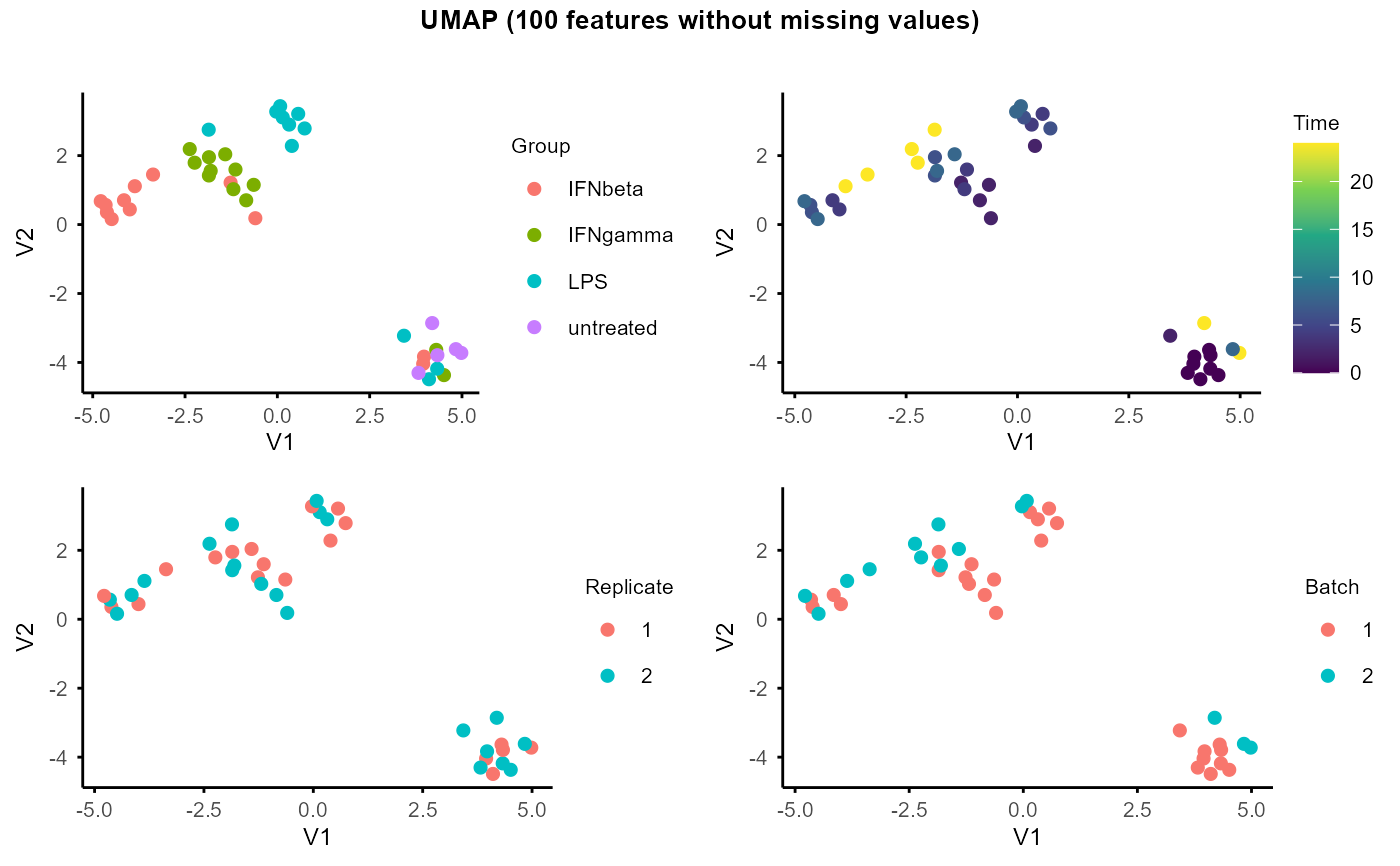 #> Warning: Using size for a discrete variable is not advised.
#> Warning: Using size for a discrete variable is not advised.